
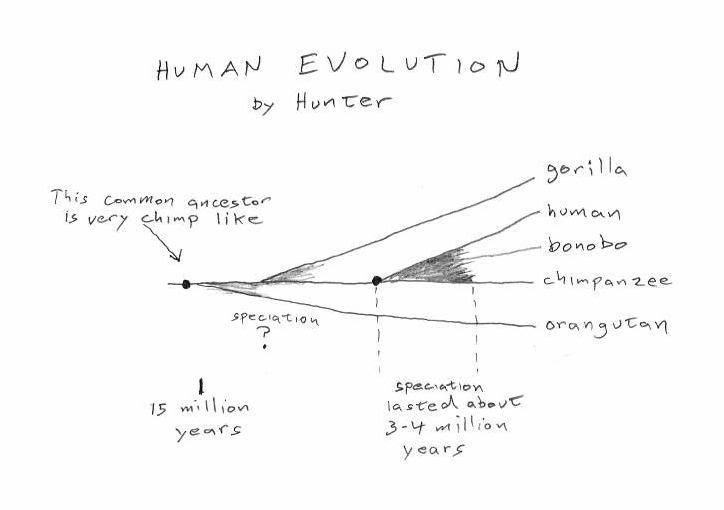
The relative age of genetic changes between the human and chimp genomes varies over a period of ~ 4 million years, depending on where in the genome you look. This figure illustrates this concept (note -- these are not actual data points). For example, Gene A first diverged millions of years before Gene B. Of particular note is the X chromosome, which falls almost entirely in the time just before final speciation. Credit: Broad Institute
New study suggests that the last common ancestor shared between chimps and humans may be ~1 million years more recent than previous estimates. Additional findings reveal a particularly young age of one of the human sex chromosomes and point to a complex process of speciation, with possible interbreeding during speciation.

This is the second family of chimpanzee endogenous retroviruses with no orthologue in the human genome. We identified ten solo LTRs and eight full-length copies of CERV 2 elements in the chimpanzee genome although, because of incomplete sequencing, we could identify the LTRs for only four of the eight full-length elements. CERV 2 elements are typically larger than CERV 1/PTERV1 elements, ranging in size from 8 to 10 kb in length. CERV 2 elements are bordered by inverted terminal repeats (TG and CA), have 4 bp TSDs (Table 2) and a proline tRNA primer binding site (Table 2). The LTRs of the CERV 2 family of elements range from 486 to 497 bp in length. Based on their LTR sequence identity (98.07% to 99.6%), we estimate that full-length CERV 2 elements were transpositionally active in the chimpanzee genome between 1.3 and 6.0 MYA. Thus, the majority of CERV 2 elements were biologically active after the divergence of chimpanzees and humans from a common ancestor.
Phylogenetic analysis of solo LTRs and LTRs from full-length elements revealed that CERV 2 elements group into at least four subfamilies (bootstrap values >95; Figure 4). We estimated the ages of two of the more abundant subfamilies by calculating the average of the pairwise distances between all sequences in each subfamily. The estimated ages of the two subfamilies were 21.9 MY and 14.1 MY, respectively. As was the case for the CERV 1/PTERV1 family, these age estimates are inconsistent with the fact that no CERV 2 orthologues were detected in the sequenced human genome. Again, we were able to detect pre-integration sites at those regions in the human genome orthologous to the CERV 2 insertion sites in chimpanzees, effectively eliminating the possibility that the elements were once present in humans but subsequently excised.
It will now be argued that a model based upon hybridisation actually meets these criticisms far better.
The model follows the lines outlined by Templeton (1981) and described in King (1993 p 240-1) with a number of small but important modifications.
His step 4 (the evolution of distinct karyotypes) has been merged into step 1 (the initial hybridisation) providing a mechanism for rapid (immediate) speciation and his steps 4 & 5 are not needed.
1. A hybridisation of two moderately long-separated (about between 1.5 and 3 my.) populations occurs, resulting in several F1 progeny born with the heterozygous form of the chromosomal fusion at the same time and at the same place. The hybrids include greater genetic variation than any born to parents both from the same populations. Most are inviable but some survive to adulthood and are fertile.
2. Several surviving heterozygous F1s interbreed producing homozygous F2 and beyond generations still with high variability from which the most fit genotypes are quickly selected for. This subpopulation is already automatically reproductively isolated from both parental populations, through the homozygous form of the chromosomal aberration.
3. Once a stable and successful recombinant form evolved it would either coexist with the parental group or expand into a new area causing a new distribution and perhaps displace the parental groups eventually.
The first criticism of the other models suggest that they offered no plausible benefit that was strong enough to overcome the accepted heterozygous disadvantage.
If the mutation occurred in a single population all a chromosomal rearrangement is essentially doing is ‘re-packaging’ old information. But if the mutation was the result of a hybridisation then at least the rearrangement is accommodating genic data from two populations, providing a great deal of variation. Out of that, potentially, will arise individuals fitter than either of the parental populations, especially if born in a new hybrid zone to which neither population are yet adapted.
According to the hybridisation model several F1 hybrids would be born at the same time and the same place. In this regard it differs greatly from most of the others in that it does not need small founding populations. Indeed the bigger the populations involved in interbreeding the more progeny with the heterozygote condition of the chromosomal rearrangement there would be.
On the face of it the case for speciation through hybrid recombination would appear to contradict the theory of chromosomal evolution. The former argues that hybrids should exhibit greater fitness than the parental population, the later that they should be less fit. However this contradiction does not arise when chronological factors are considered.
See Fig 4 (below) to help with the following argument that hybrid viability will logically vary with time.
 In a, one population with 48 chromosomes splits into two which are then reproductively isolated from each other and continue to be so. After sufficient time has elapsed enough genic changes and mild chromosomal rearrangements (such as small translocations, insertions and deletions but not chromosomal fusions or inversions) have arisen to create a genetic barrier to hybridisation.
In a, one population with 48 chromosomes splits into two which are then reproductively isolated from each other and continue to be so. After sufficient time has elapsed enough genic changes and mild chromosomal rearrangements (such as small translocations, insertions and deletions but not chromosomal fusions or inversions) have arisen to create a genetic barrier to hybridisation.
In b, two populations become reproductively isolated, but only briefly. Genetic drift began but not enough to cause any problems when the populations re-merged. The population remains one species although genetic drift continues to change its phenotype over time.
The situation in c, is intermediate between the other two. The populations are separated for long enough for some genetic barriers (for instance a small translocation or deletion) to have begun to block the gene flow, but not sufficient to stop it completely. Instead, when they re-merge a variety of hybrids form at the same time and place with significant chromosomal rearrangements. The fittest of these survive and begin a new species (in the strict sense) with a different chromosome number.The Hybrid Zone factor
A similar argument could be made in terms of the ecology of the two populations. If one coherent, constantly interbreeding population lived in the same place the matter of speciation simply would not arise. If two species become adapted to two very different and geographically isolated niches the hybrids might not be viable at all but if the two populations lived in two similar, but different niches, separated by a hybrid zone one could easily see how a hybrid might actually be more fit that its parental groups.
As Arnold (1999 p 372) put it “hybrid genotypes can have high fitness compared to parental genotypes and even if hybrids demonstrate low fitness, they can still act as the starting point for new evolutionary lineages.”
Hybrid zones have been known to give rise to so called ‘hybrizymes’ or ‘rare alleles’ that are not present in the parental populations (Schilthuizen et al. 1999 p 2181.) Their field study, of the hybrid zone between two Greek land snails Albinaria hippolyti holtzi and A. h. aphrodite, found that a haplotype of an allele that was rare in A. h. Aphrodite and totally absent from A. h. holtzi was quite common in the hybrid zone.
The ‘hybrid dysgenesis’ phenomena (King 1993 p 256) where crossed species of Drosophila have been observed to produce unusual effects in hybrids produced the laboratory only adds more weight to the view that hybrid zones can increase genetic variation. It is logical that from out of this extra variation some of the genotypes may even be fitter than their parental populations.
When Baird et al (2000) published their paper proposing “a model that involves the hybridisation of two archaic hominoid lineages ultimately giving rise to Homo sapiens”, they were the first to do so, as far as they or I know.
Their evidence for this has nothing to do with chromosome numbers or, in fact, any chromosomal aberration at all. Their conclusion comes from a quite different and seemingly far more complex area of study: sequence polymorphism and linkage disequilibrium at the telomere junction on human chromosomes Xp and 12q.
Basically, in studying sequences of telomere-adjacent sequences of the human genome they found relatively few haplotypes of which two main ones were highly diverged. They assumed that the regions were well conserved. (As they put it “provided that the telomere-adjacent sequences in the genome remain recombinationally suppressed, diverged haplotypes that arise in these regions may persist for a very long time.”) So they tried to come up with a scenario that might explain the observation.
They concluded that the most plausible explanation was that the haplotypes (they called ‘B’ and ‘D’) arose “in separate archaic hominoid lineages, from a common ancestral sequence… separated for sufficient time to allow divergent haplotypes to arise as a result of sequential mutations and of fixation in each lineage of one predominant haplotypes.”
They even went on to attempt to calculate the length of time the lineages were separated. This is theoretically possible by calculating the sequence divergence between types B and D in humans and between types B in humans and chimpanzees and multiplying the ratio of the two by the estimated point of coalescence between humans and chimpanzees.
The calculation they came up with was therefore…
BPan-BHomo (4.6%) /B-Dhomo (1.9%) ? Pan/Homo lca (4.5 mya) = 1.9 mya.
Their figure of 4.5 million years for the Pan/Homo split may be in doubt (especially in the light of the discovery of the potentially bipedal hominid Orrorin dated at 6 mya) but the figure was really only meant as an estimate.
Finally, it should be mentioned that in a personal correspondence with one of the authors of the paper (N. J. Royle from Leicester University) it was made clear that they do not see differences in chromosome number as evidence for a hominid hybridisation event.
Assuming that at least one of the three major chromosomal rearrangements observed in the human karyotype (the telomeric fusion in chromosome 2, the pericentric inversion or the reciprocal translocation on chromosome 5) was the result of a hybridisation event of two ancestral hominoid species we can turn to the exciting area of speculation as to when and where this might have happened.
Using Baird et al.’s (2000 p 248) data and calculation, but assuming a last common ancestor of Pan/Homo of 5.5 million years (rather than their 4.5 million years), a figure of 2.3 million years can be calculated as the length of time the two lineages were separated.
This gives us a simple sliding scale against which to postulate when the hybridisation event occurred and which paleospecies may have been responsible.
The fig below uses the Hominid phylogenetic tree and timescale printed in Klein (1999 p 227) as a template in order to estimate when the hybridisation event might best fit.

It is clear from this that the proposed hybridisation could theoretically have occurred at any time between 3.7 million years ago and 200,000 ago (the assumed latest date for the emergence of Homo sapiens from Klein 1999 p 507.) According to this, any (apart from Australopithecus afarensis) of our putative ancestors could have been the result of a hybridisation.
There is enough time, for instance that Australopithecus africanus resulted from the hybridisation of two ape-like ancestors, although there are very few fossil candidates to have participated in such a hybridisation.
A much more compelling case is that Homo sapiens is, in fact, the result of the hybridisation. Assuming that the first anatomically modern human lived between 500,000 and 200,000 years ago this would open up a window allowing practically any of the Homo or Paranthropus species in our phylogeny to be seen as candidates for such a hybridisation.
Assuming that Homo sapiens is indeed the result of a hybridisation of two hominids, we may conclude that the speciation event most likely occurred around 500,000 years ago in East Africa between two hominid species that had been separated for over 2 million years.
From our studies of chromosomal rearrangements we may speculate that the two hominids, say Homo ergaster and perhaps another as yet un-discovered hominid, both had 48 chromosomes, as did all the hominids in our phylogeny.
It would logically follow from this that of all the different hominids that have ever lived only Homo sapiens evolved a significant postzygotic reproductive barrier preventing successful cross-breeding with other types.
If true, this would have quite a bearing on our understanding of human evolution. It would, for instance, shed new light on ambiguities in the fossil record, where two paleospecies appear to have traits suggesting conflicting phylogenies. (For example the placement of Paranthropus aethiopicus - see Klein pp 222 - 229 for discussion.) Perhaps the so-called “muddle-in-the-middle” is not muddle at all, but simply evidence for several hybridisation events that occurred between hominid groups 3 and 2 million years ago that were not our direct ancestors..
Furthermore it would add significant weight behind the “Out-of-Africa II theory” or the “replacement model,” which suggests that Homo sapiens emerged in Africa quite recently (about 150,000 years ago) and then spread out across the world replacing all the other hominids without any significant interbreeding. If the hybridisation model is right and the change from 48 to 46 chromosomes introduced a novel and relatively impenetrable genetic barrier it would surely explain how so little interbreeding seems to have taken place according to the genetic and molecular data.
The hybridisation model may even shed some light on areas of human evolution that have hitherto proved very difficult to explain but perhaps it would be best to leave that speculation for a later date.
7. Questions & Ideas for further study
This study has been tremendously enjoyable and stimulating for me personally. Once I got into the subject I found that it drew me in further. The more I learnt the more ignorant I felt but, at the same time, the more evidence there seemed to be for the hybridisation idea I had originally started out with.
The study only had a finite time to run and so I’ve had to finish here. I’d finally just like to note down some ideas of unanswered questions and ideas for further study.
· How does chromosomal evolution affect the phylogenetic relationships and estimates of last common ancestor from the ‘molecular clock?
If the model proved correct, what impact (if any) would this have on the currently accepted dates that have been calculated by the molecular clock? Do the figures currently account for whether a population may have interbred with external populations? For instance, if it turns out that Homo sapiens has not interbred with any other hominoids in the last 200,000 years but that Pan had done so many times, would this change our assumed dates in any way?
· Do gorillas and chimpanzees produce viable hybrids?
I was not able to find any evidence of Pan ? Gorilla hybridisation although Sommer (2000 pers. comment) suggested that it had been known although he did not known about the fertility of the offspring. Such studied, although probably unethical, would shed some light onto this debate. The reciprocal translocation on Gorilla chromosome 5 would, according to this model, be sufficient to prevent a viable hybrid but it would be interesting to know. Evidence into other crosses, even more unethical, was also absent.
· Did neanderthals have 48 chromosomes or 46?
Another issue thrown up by this debate is whether neanderthals had 48 or 46 chromosomes. According to the model described here, it would seem likely that they also had 48. Perhaps they too resulted from an African hominid hybridisation but that because it occurred another 500,000 years earlier the full chromosomal rearrangement did not occur with them.
· Can similar results produced by Baird et al. (2000) be reproduced on Y chromosome studies?
It would be interesting to see if there were any similar groupings of conservative micro-satellites as they found near the telomere junctions of autosomes. If so, it would add support to the theory.
· Can telomere-telomere fusions occur during fertilization as well as during meiosis I?
It occurred to me during this study that chromosomal rearrangements are only thought happen in the early phases of meiosis. Is it not also possible that they may occur during fertilization of the gametes? It would seem plausible that telomere-telomere fusions, in particular, could occur at this late stage or perhaps the mitotic divisions immediately after fertilization. They seem to be implicated, for instance, in certain kinds of tumours - a purely mitotic division. If so, this would make the idea of the immediate rearrangement of the initial hybrid much more plausible.